by
John R. Fischer, Senior Reporter | July 06, 2023
A new approach that combines genomics and PET tau imaging data could potentially personalize treatment for Alzheimer's disease.
Alzheimer’s disease is divided into different subtypes that vary in symptoms and prognosis. In the past, researchers depended on a single model to describe the development of tau protein tangles, one of the hallmarks of the disease, despite there being cases that did not align with this model.
In Massachusetts, scientists recently applied a new computational approach combining genomics and tau PET imaging data that identified four types of AD and genes most associated with each. The approach is based on a novel clustering framework that uses sparse canonical correlation analysis (SCCA), where two sets of variables that highly correlate with one another are combined.
They discussed their findings at the 2023 Society of Nuclear Medicine and Molecular Imaging Annual Meeting, saying that with more research, their technique may provide insights that could potentially personalize treatment of subtypes in the future.



Ad Statistics
Times Displayed: 5361
Times Visited: 18 Brand-New FDA-cleared Advanced Ultrasound Medical Device available for sale or lease to Wound Care Centers or any other Medical Facilities.The Arobella 1000D is designed for non-contact or debridement ultrasound wound healing therapy, or any other wounds
“Genomics- and imaging-guided individualized subtyping is vital for Alzheimer’s disease because different subtypes may also have distinct rates and profiles of cognitive decline, potentially affecting clinical trial outcomes and treatment response,” said Joyita Dutta, associate professor in the department of biomedical engineering at the University of Massachusetts in Amherst, in a statement.
For their study, the researchers used tau imaging data with uptake value ratios from 10 broad regions in 334 cognitively normal and 207 cognitively impaired patients who underwent 18F-flortaucipir PET scans. They also used Illumina SNP genotyping data from the patients, which identified and extracted 145 genome variations associated with AD.
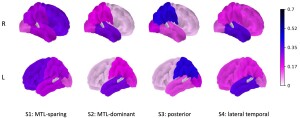
The SCCA-clustering framework was jointly applied to the tau PET and genomics data sets, identifying four subtypes: medial temporal lobe (MTL)-dominant, posterior, MTL-sparing, and lateral-temporal. Along with the APOE gene, it also showed correlations between top genes and each subtype, including ADAMTS4 for the MTL-dominant subtype, TRPM1 for posterior, CYP4V2 for MTL-sparing, and CR1 for lateral-temporal subtype.
The researchers say that the approach could also be applied to many other types of diseases.
Imaging and genomics data came from the Alzheimer's Disease Neuroimaging Initiative (ADNI).

