by
John R. Fischer, Senior Reporter | December 15, 2020
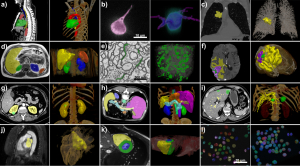
The technique enables users with little specialist knowledge to configure self-learning algorithms
The German Cancer Research Center (DKFZ) has a new method for generating self-learning algorithms from large numbers of diverse imaging data sets.
Known as nnU-Net, the technique is expected to allow users with little to no specialist knowledge configure these self-learning tools for specific tasks.
"nnU-Net can be used immediately, can be trained using imaging data sets, and can perform special tasks — without requiring any special expertise in computer science or any particularly significant computing power," said study director professor Dr. Klaus Maier-Hein of DKFZ’s division of medical image computing.



Ad Statistics
Times Displayed: 138115
Times Visited: 7975 MIT labs, experts in Multi-Vendor component level repair of: MRI Coils, RF amplifiers, Gradient Amplifiers Contrast Media Injectors. System repairs, sub-assembly repairs, component level repairs, refurbish/calibrate. info@mitlabsusa.com/+1 (305) 470-8013
In algorithm development, computers must learn to interpret three-dimensional imaging data sets, and how to differentiate between tumor and non-tumor pixels. Imaging data sets where tumors, healthy tissue and other anatomical structures have been labeled by hand are used as training materials.
nnU-Net adapts dynamically and automatically to any kind of imaging data set to allow researchers with limited knowledge to configure self-learning algorithms. In addition to forming algorithms from data for modalities like MR and CT, nnU-Net processes images from electron and fluorescence microscopy. The method has obtained best results in 33 of 53 different segmentation tasks in international competitions against highly specific algorithms, according to DKFZ researchers.
Maier-Hein and his team are making it available as an open source tool that can be downloaded free of charge. While AI assessments of medical imaging data are primarily used for research, they expect their technique to reduce highly repetitive tasks in large-scale clinical studies.
"nnU-Net can help harness this potential," said Maier-Hein.
Research on the technique was published in
Nature Methods.

